rthin
rthin applies independent random thinning to a point pattern.
Description
This function was ported to matlab from the R spatstat package, developed by Adrian Baddeley (Adrian.Baddeley@curtin.edu.au), Rolf Turner (r.turner@auckland.ac.nz) and Ege Rubak (rubak@math.aau.dk) for the statistical analysis of spatial point patterns. The algorithm for random thinning was changed in spatstat version 1.42-3. Our matlab porting is based on a earlier version. See the rthin documentation in spatstat for more details.
In a random thinning operation, each point of X is randomly either deleted or retained (i.e. not deleted). The result is a point pattern, consisting of those points of X that were retained. Independent random thinning means that the retention/deletion of each point is independent of other points.
Examples
Input Arguments
Output Arguments
References
Bowman, A.W. and Azzalini, A. (1997), "Applied Smoothing Techniques for Data Analysis", Oxford University Press.
Acknowledgements
This function was ported to matlab from the R spatstat package, developed by Adrian Baddeley (Adrian.Baddeley@curtin.edu.au), Rolf Turner (r.turner@auckland.ac.nz) and Ege Rubak (rubak@math.aau.dk) for the statistical analysis of spatial point patterns. The algorithm for random thinning was changed in spatstat version 1.42-3. Our matlab porting is based on a previous version. See the rthin documentation in spatstat for more details.
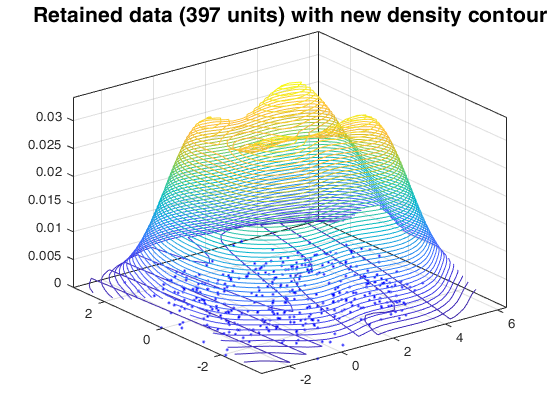
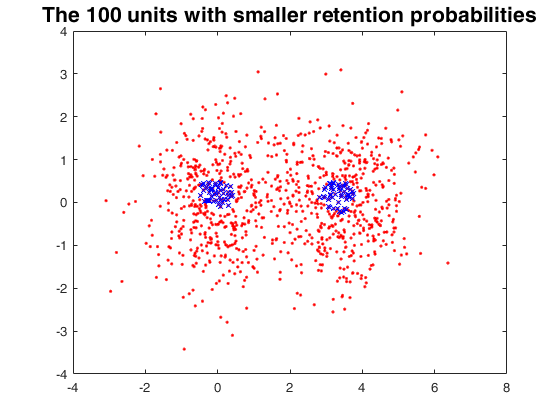
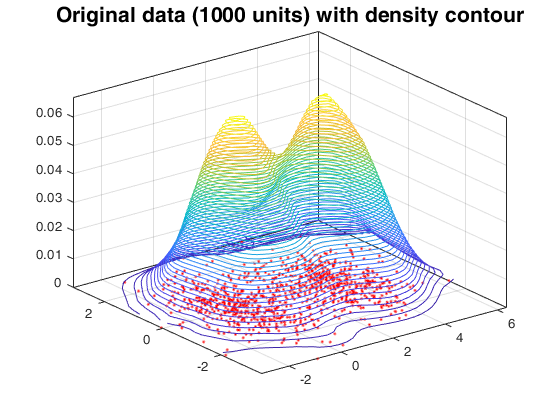
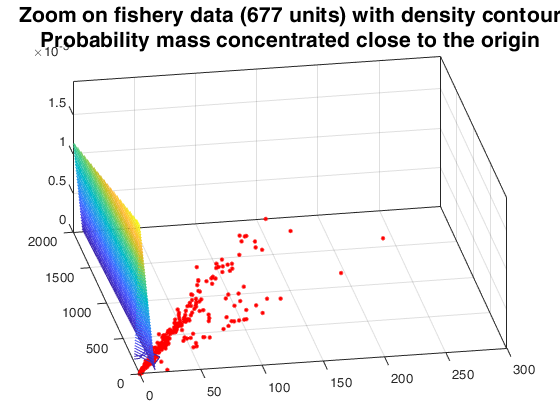
 Random thinning on a mixture of normal distribution.
Random thinning on a mixture of normal distribution.