FSMenvmmd
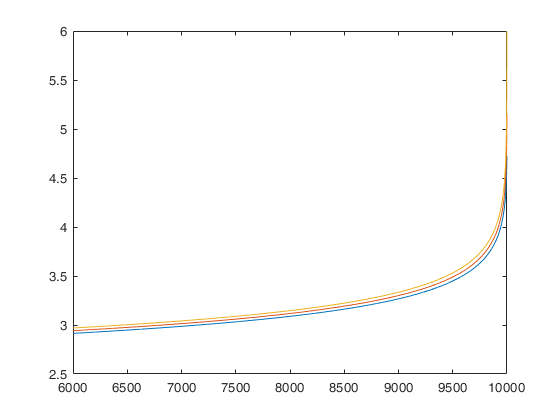
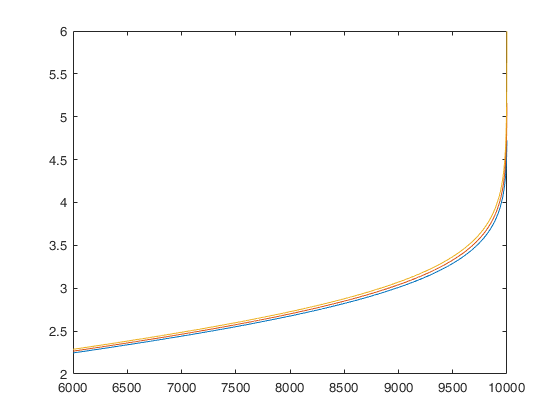
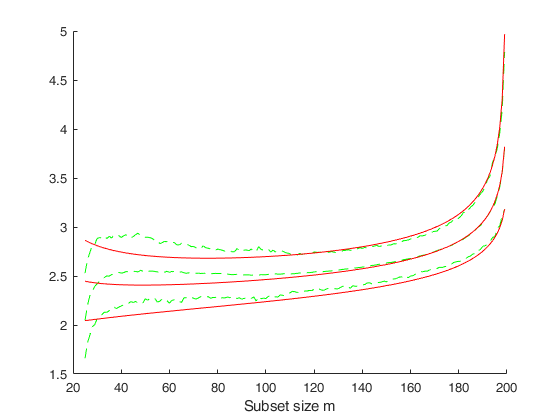
FSMenvmmd computes the theoretical envelopes of Minimum MD outside subset during the search
Description
Examples
Related Examples
Input Arguments
Output Arguments
References
Riani, M., Atkinson, A.C. and Cerioli, A. (2009), Finding an unknown number of multivariate outliers, "Journal of the Royal Statistical Society Series B", Vol. 71, pp. 201-221.
 FSMenvmmd with all default options.
FSMenvmmd with all default options.