ClusterRelabel
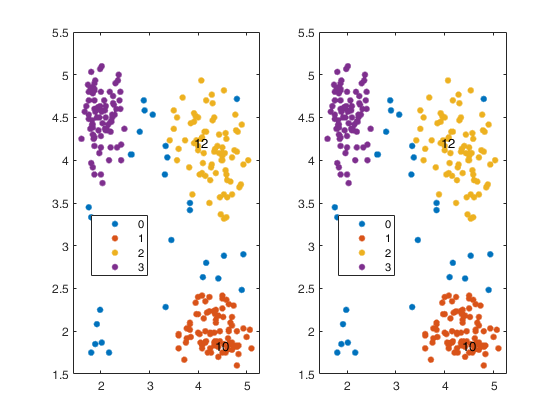
ClusterRelabel enables to control the labels of the clusters which contain predefined units
Syntax
Description
Start with labelling produced by tclustIC and produce consistent labels.IDXrelabelled
=ClusterRelabel(IDX,
pivotunits)
[
Example with detailed description of output element OldAndNewIndexes.IDXrelabelled,
idxMapping]
=ClusterRelabel(___)
Examples
Input Arguments
Output Arguments
References
Cerioli, A., Garcia-Escudero, L.A., Mayo-Iscar, A. and Riani M. (2017), Finding the Number of Groups in Model-Based Clustering via Constrained Likelihoods, "Journal of Computational and Graphical Statistics", pp. 404-416, https://doi.org/10.1080/10618600.2017.1390469
 Example with detailed description of output element OldAndNewIndexes.
Example with detailed description of output element OldAndNewIndexes.