FSRHmdr
FSRHmdr computes minimum deletion residual and other basic linear regression quantities in each step of the heteroskedastic search
Syntax
mdr=FSRHmdr(y,X,Z,bsb)examplemdr=FSRHmdr(y,X,Z,bsb,Name,Value)example[mdr,Un]=FSRHmdr(___)example[mdr,Un,BB]=FSRHmdr(___)example[mdr,Un,BB,Bgls]=FSRHmdr(___)example[mdr,Un,BB,Bgls,S2]=FSRHmdr(___)example[mdr,Un,BB,Bgls,S2,Hetero]=FSRHmdr(___)example[mdr,Un,BB,Bgls,S2,Hetero,WEI]=FSRHmdr(___)example
Description
Examples
 FSRHmdr with all default options.
FSRHmdr with all default options.
 FSRHmdr with all default options.
FSRHmdr with all default options.Common part to all examples: load tradeH dataset (used in the paper ART).
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
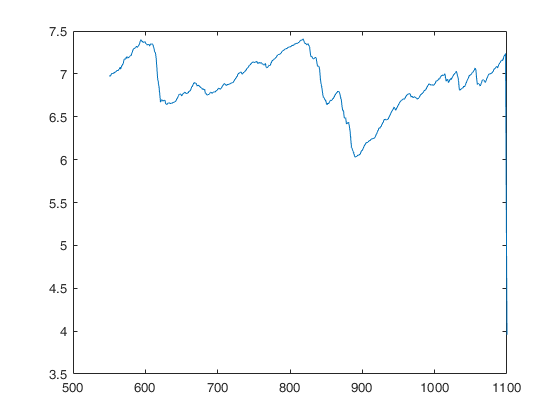
mdr=FSRHmdr(y,X,Z,0);
Warning: interchange greater than 10 when m=22 Number of units which entered=21 Warning: interchange greater than 10 when m=24 Number of units which entered=14 Warning: Matrix is singular, close to singular or badly scaled. Results may be inaccurate. RCOND = NaN. Warning: interchange greater than 10 when m=25 Number of units which entered=24 Warning: interchange greater than 10 when m=26 Number of units which entered=27 Warning: interchange greater than 10 when m=165 Number of units which entered=11 Warning: interchange greater than 10 when m=166 Number of units which entered=39 Warning: interchange greater than 10 when m=167 Number of units which entered=11 Warning: interchange greater than 10 when m=168 Number of units which entered=15 Warning: interchange greater than 10 when m=169 Number of units which entered=16 Warning: interchange greater than 10 when m=170 Number of units which entered=13 Warning: interchange greater than 10 when m=171 Number of units which entered=11 Warning: interchange greater than 10 when m=172 Number of units which entered=11 Warning: interchange greater than 10 when m=180 Number of units which entered=12 Warning: interchange greater than 10 when m=181 Number of units which entered=25 Warning: interchange greater than 10 when m=182 Number of units which entered=15 Warning: interchange greater than 10 when m=183 Number of units which entered=18 Warning: interchange greater than 10 when m=225 Number of units which entered=15 Warning: interchange greater than 10 when m=482 Number of units which entered=26 Warning: interchange greater than 10 when m=483 Number of units which entered=11 Warning: interchange greater than 10 when m=484 Number of units which entered=12 Warning: interchange greater than 10 when m=485 Number of units which entered=12 Warning: interchange greater than 10 when m=486 Number of units which entered=11
FSRHmdr with optional arguments.
Specifying the search initialization.
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
mdr=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
Analyze units entering the search in the final steps.
Compute minimum deletion residual and analyze the units entering subset in each step of the fwd search (matrix Un). As is well known, the FS provides an ordering of the data from those most in agreement with a suggested model (which enter the first steps) to those least in agreement with it (which are included in the final steps).
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
[mdr,Un]=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
Units forming subset in each step.
Obtain detailed information about the units forming subset in each step of the forward search (matrix BB).
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
[mdr,Un,BB]=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
Monitor \hat \beta.
Monitor how the estimates of beta coefficients changes as the subset size increases (matrix Bols).
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
[mdr,Un,BB,Bols]=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
Monitor s^2.
Monitor the estimate of sigma^2 in each step of the fwd search (matrix S2).
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
[mdr,Un,BB,Bols,S2]=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
Monitoring the estimates of the weights.
XX=load('tradeH.txt');
y=XX(:,2);
X=XX(:,1);
X=X./max(X);
Z=log(X);
[mdr,Un,BB,Bols,S2,Hetero,WEI]=FSRHmdr(y,X,Z,0,'init',round(length(y)/2));
plot(S2(:,1),WEI')
title('Monitoring of the weights')
Input Arguments
y — Response variable.
Vector.
Response variable, specified as a vector of length n, where n is the number of observations. Each entry in y is the response for the corresponding row of X.
Missing values (NaN's) and infinite values (Inf's) are allowed, since observations (rows) with missing or infinite values will automatically be excluded from the computations.
Data Types: single| double
X — Predictor variables in the regression equation.
Matrix.
Matrix of explanatory variables (also called 'regressors') of dimension n x (p-1) where p denotes the number of explanatory variables including the intercept.
Rows of X represent observations, and columns represent variables. By default, there is a constant term in the model, unless you explicitly remove it using input option intercept, so do not include a column of 1s in X. Missing values (NaN's) and infinite values (Inf's) are allowed, since observations (rows) with missing or infinite values will automatically be excluded from the computations.
Data Types: single| double
Z — Predictor variables in the scedastic equation.
Matrix.
n x r matrix or vector of length r.
If Z is a n x r matrix it contains the r variables which form the scedastic function as follows (if input option art==1)
\omega_i = 1 + exp(\gamma_0 + \gamma_1 Z(i,1) + ...+ \gamma_{r} Z(i,r)) If Z is a vector of length r it contains the indexes of the columns of matrix X which form the scedastic function as follows \omega_i = 1 + exp(\gamma_0 + \gamma_1 X(i,Z(1)) + ...+ \gamma_{r} X(i,Z(r))) Therefore, if, for example, the explanatory variables responsible for heteroscedasticity are columns 3 and 5 of matrix X, it is possible to use both the syntax FSRHmdr(y,X,X(:,[3 5]),0) or the syntax FSRHmdr(y,X,[3 5],0)
Data Types: single| double
bsb — list of units forming the initial subset.
Vector | 0.
If bsb=0 then the procedure starts with p units randomly chosen else if bsb is not 0 the search will start with m0=length(bsb)
Data Types: single| double
Name-Value Pair Arguments
Specify optional comma-separated pairs of Name,Value arguments.
Name is the argument name and Value
is the corresponding value. Name must appear
inside single quotes (' ').
You can specify several name and value pair arguments in any order as
Name1,Value1,...,NameN,ValueN.
'init',100 starts monitoring from step m=100
, 'intercept',false
, 'typeH','har'
, 'plots',1
, 'nocheck',true
, 'msg',1
, 'gridsearch',0
, 'constr',[1 6 3]
, 'bsbmfullrank',0
,
init
—Search initialization.scalar.
It specifies the point where to start monitoring required diagnostics. If it is not specified it is set equal to: p+1, if the sample size is smaller than 40;
min(3*p+1,floor(0.5*(n+p+1))), otherwise.
The minimum value of init is 0. In this case, in the first step we just use prior information
Example: 'init',100 starts monitoring from step m=100
Data Types: double
intercept
—Indicator for constant term.true (default) | false.
Indicator for the constant term (intercept) in the fit, specified as the comma-separated pair consisting of 'Intercept' and either true to include or false to remove the constant term from the model.
Example: 'intercept',false
Data Types: boolean
typeH
—Parametric function to be used in the skedastic equation.string | character.
If typeH is 'art' (default) than the skedastic function is modelled as follows \sigma^2_i = \sigma^2 (1 + \exp(\gamma_0 + \gamma_1 Z(i,1) + \cdots + \gamma_{r} Z(i,r))) on the other hand, if typeH is 'har' then traditional formulation due to Harvey is used as follows \sigma^2_i = \exp(\gamma_0 + \gamma_1 Z(i,1) + \cdots + \gamma_{r} Z(i,r)) =\sigma^2 (\exp(\gamma_1 Z(i,1) + \cdots + \gamma_{r} Z(i,r))
Example: 'typeH','har'
Data Types: character or string
plots
—Plot on the screen.scalar.
If equal to one a plot of Bayesian minimum deletion residual appears on the screen with 1 per cent, 50 per cent and 99 per cent confidence bands; else (default) no plot is shown.
Remark. The plot which is produced is very simple. In order to control a series of options in this plot and in order to connect it dynamically to the other forward plots, it is necessary to use function mdrplot
Example: 'plots',1
Data Types: double
nocheck
—Check input arguments.boolean.
If nocheck is equal to true, no check is performed on matrix y and matrix X. Notice that y and X are left unchanged. In other words, the additional column of ones for the intercept is not added. As default nocheck=false.
Example: 'nocheck',true
Data Types: boolean
msg
—Level of output to display.scalar.
It controls whether to display or not messages about great interchange on the screen.
If msg==1 (default), messages are displayed on the screen, else no message is displayed on the screen.
Example: 'msg',1
Data Types: double
gridsearch
—Algorithm to be used.scalar.
If gridsearch ==1, grid search will be used, else the scoring algorithm will be used.
REMARK: the grid search has only been implemented when there is just one explanatory variable which controls heteroskedasticity.
Example: 'gridsearch',0
Data Types: double
constr
—units which are forced to join the search in the last r steps.vector.
r x 1 vector. The default is constr=''. No constraint is imposed
Example: 'constr',[1 6 3]
Data Types: double
bsbmfullrank
—It tells how to behave in case subset at step m
(say bsbm) produces a non singular X.scalar.
In other words, this options controls what to do when rank(X(bsbm,:)) is smaller than the number of explanatory variables. If bsbmfullrank = 1 (default is 1), these units (whose number is say mnofullrank) are constrained to enter the search in the final n-mnofullrank steps, else the search continues using as an estimate of beta at step m the estimate of beta found in the previous step.
Example: 'bsbmfullrank',0
Data Types: double
bsbsteps
—Save the units forming subsets (and weights vector) in
selected steps.vector.
It specifies for which steps of the fwd search it is necessary to save the units forming subset. If bsbsteps is 0, we store the units forming subset in all steps. The default is to store the units forming subset in all steps if n<=5000, otherwise store the units forming subset at steps init and steps that are multiples of 100. For example, as default, if n=7530 and init=6, units forming subset are stored for m=init, 100, 200, ..., 7500.
Example: 'bsbsteps',[100 200] stores the unis forming
subset in steps 100 and 200.
Data Types: double
Output Arguments
mdr —Minimum deletion residual.
Matrix
n -init x 2 matrix which contains the monitoring of minimum deletion residual at each step of the forward search.
1st col = fwd search index (from init to n-1).
2nd col = minimum deletion residual.
REMARK: if in a certain step of the search matrix is singular, this procedure checks how many observations produce a singular matrix. In this case, mdr is a column vector which contains the list of units for which matrix X is non singular.
Un —Units included in each step.
Matrix
(n-init) x 11 Matrix which contains the unit(s) included in the subset at each step of the search.
REMARK: in every step the new subset is compared with the old subset. Un contains the unit(s) present in the new subset, but not in the old one.
Un(1,2), for example, contains the unit included in step init+1.
Un(end,2) contains the units included in the final step of the search.
BB —Units belonging to subset in each step.
Matrix
n x (n-init+1) or n-by-length(bsbsteps) matrix (depending on input option bsbsteps) which contains information about the units belonging to the subset at each step of the forward search.
BB has the following structure 1-st row has number 1 in correspondence of the steps in which unit 1 is included inside subset and a missing value for the other steps ......
(n-1)-th row has number n-1 in correspondence of the steps in which unit n-1 is included inside subset and a missing value for the other steps n-th row has number n in correspondence of the steps in which unit n is included inside subset and a missing value for the other steps.
The size of matrix BB is n x (n-init+1), if option input bsbsteps is 0, else the size is n-by-length(bsbsteps).
Bgls —GLS beta coefficients.
Matrix
(n-init+1) x (p+1) matrix containing the monitoring of estimated beta coefficients in each step of the forward search.
S2 —S2 and R2.
Matrix
(n-init+1) x 3 matrix containing the monitoring of S2 (2nd column)and R2 (third column) in each step of the forward search.
Hetero —Coefficients in the heteroskedastic equation.
Matrix
(n-init1+1) x (r+1) matrix containing: 1st col = fwd search index;
2nd col = estimate of first coeff in the scedastic equation;
...
(r+1)-th col = estimate of last coeff in the scedastic equation.
WEI —Weights.
Matrix
n x (n-init+1) or n-by-length(bsbsteps) matrix (depending on input option bsbsteps) which contains information about the weights assigned to each unit to make the regression equation skedastic.
More precisely, if var (\epsilon)= \sigma^2 Omega=diag(omegahat) the weights which are stored are omegahat.^(-0.5);
References
Atkinson, A.C., Riani, M. and Torti, F. (2016), Robust methods for heteroskedastic regression, "Computational Statistics and Data Analysis", Vol. 104, pp. 209-222, http://dx.doi.org/10.1016/j.csda.2016.07.002 [ART]