FSCorAnaeda
FSCorAnaeda performs forward search in correspondence analysis with exploratory data analysis purposes
Description
Examples
FSCorAnaeda with all default options.
Generate contingency table of size 50-by-5 with total sum of n_ij=2000.
I=50; J=5; n=2000; % nrowt = column vector containing row marginal totals nrowt=(n/I)*ones(I,1); % ncolt = row vector containing column marginal totals ncolt=(n/J)*ones(1,J); out1=rcontFS(I,J,nrowt,ncolt); N=out1.m144; RAW=mcdCorAna(N,'plots',0); ini=round(sum(sum(RAW.N))/4); out=FSCorAnaeda(RAW);
 FSCorAnaeda with optional arguments.
FSCorAnaeda with optional arguments.
 FSCorAnaeda with optional arguments.
FSCorAnaeda with optional arguments.Generate contingency table of size 50-by-5 with total sum of n_ij=2000.
I=50; J=5; n=2000; % nrowt = column vector containing row marginal totals nrowt=(n/I)*ones(I,1); % ncolt = row vector containing column marginal totals ncolt=(n/J)*ones(1,J); out1=rcontFS(I,J,nrowt,ncolt); N=out1.m144; RAW=mcdCorAna(N,'plots',0); ini=round(sum(sum(RAW.N))/4); out=FSCorAnaeda(RAW,'plots',1);
Total estimated time to complete MCD: 0.04 seconds Creating empirical confidence band for minimum (weighted) Mahalanobis distance

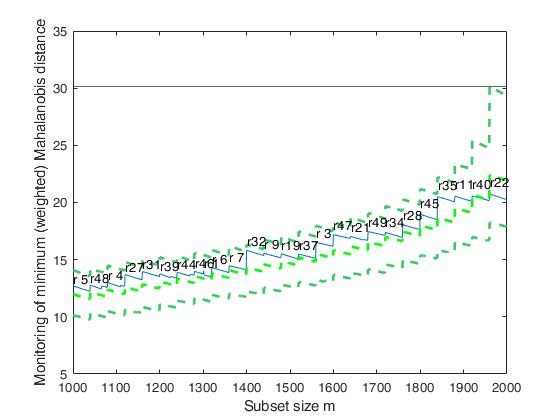
Related Examples
FSCorAnaeda starting from a random initial subset.
Generate contingency table of size 50-by-5 with total sum of n_ij=2000.
I=50; J=5; n=2000; % nrowt = column vector containing row marginal totals nrowt=(n/I)*ones(I,1); % ncolt = row vector containing column marginal totals ncolt=(n/J)*ones(1,J); out1=rcontFS(I,J,nrowt,ncolt); N=out1.m144; % The first input argument is a contingency table and no initial subset % and no initial location is supplied out=FSCorAnaeda(N,'plots',1);
Input Arguments
N — contingency table or structure.
Array or table of size I-by-J or structure.
If N is a structure it contains the following fields:
| Value | Description |
|---|---|
N |
contingency table in array format of size I-by-J. |
Ntable |
contingency table in table format of size I-by-J. |
loc |
initial location estimate for the matrix of Profile rows of the contingency table (row vector or length J). |
weights |
I x 1 vector containing the proportion of the mass of each rows of matrix N in the computation of the MCD estimate of location. If N.weigths(2)=0.1 it means that row 2 of the contingency table contributes with 10 per cent of its mass. The initial subset is based on N.weights. |
NsimStore |
array of size I-by-J times nsimul containing in each column the nsimul simulated contingency tables. Note that input structure N can be conveniently created by function mcdCorAna. If N is not a struct it is possible to specify the rows of the contingency table forming initial subset with input option bsb. |
Data Types: single|double
Name-Value Pair Arguments
Specify optional comma-separated pairs of Name,Value arguments.
Name is the argument name and Value
is the corresponding value. Name must appear
inside single quotes (' ').
You can specify several name and value pair arguments in any order as
Name1,Value1,...,NameN,ValueN.
'bsb',[3 6 8 10 12 14]
, 'conflev',0.99
, 'init',50
, 'plots',0
, 'msg',0
bsb
—Initial subset.vector of positive integers containing the indexes of the rows of the contingency table which have to be used to initialize the forward search.
If bsb is empty and required input argument is a struct N.loc will be used.
If bsb is supplied and N is a struct N.loc is ignored.
The default value of bsb is empty, and if N is not a struct a random subset containing round(n/5) units will be used.
Example: 'bsb',[3 6 8 10 12 14]
Data Types: double
conflev
—simultaneous confidence interval to declare units as
outliers.scalar.
The default value of conflev is 0.99, that is a 99 per cent simultaneous confidence level.
Confidence level are based on simulated contingency tables. This input argument is ignored if optional input argument mmdEnv is not missing
Example: 'conflev',0.99
Data Types: numeric
init
—Point where to start monitoring required diagnostics.scalar.
Note that if init is not specified it will be set equal to floor(n*0.6).
where the total number of units in the contingency table.
Example: 'init',50
Data Types: double
plots
—It specify whether it is necessary to produce the plots of the
monitoring of minMD.scalar.
If plots=1, a plot of the monitoring of minMD among the units not belonging to the subset is produced on the screen with 1 per cent, 50 per cent and 99 per cent confidence bands else (default), all plots are suppressed.
Example: 'plots',0
Data Types: double
msg
—It controls whether to display or not messages
about great interchange on the screen.scalar.
If msg==1 (default) messages are displayed on the screen else no message is displayed on the screen.
Example: 'msg',0
Data Types: double
Output Arguments
out — description
Structure
Structure which contains the following fields
| Value | Description |
|---|---|
MAL |
I x (n-init+1) = matrix containing the monitoring of Mahalanobis distances. 1st row = distance for first row; ...; Ith row = distance for Ith row. |
BB |
I-by-(n-init+1) matrix containing the information about the units belonging to the subset at each step of the forward search. 1st col = indexes of the units forming subset in the initial step; ...; last column = units forming subset in the final step (all units). Note that the numbers inside out.BB vary in the interval [0 1] and represent the proportions in which each unit is represented in the subset. 0.1 means that the associated row is represented in the subset with 10 per cent of its mass. |
mmd |
n-init-by-2 matrix which contains the monitoring of minimum MD or (m+1)th ordered MD at each step of the forward search. 1st col = fwd search index (from init to n-1); 2nd col = minimum MD; |
ine |
n-init-by-2 matrix which contains the monitoring of inertia at each step of the forward search. 1st col = fwd search index (from init to n); 2nd col = inertia; |
Loc |
(n-init+1)-by-J matrix containing the monitoring of estimated means for each variable in each step of the forward search. |
Un |
(n-init) x 11 Matrix which contains the unit(s) included in the subset at each step of the fwd search. REMARK: in every step the new subset is compared with the old subset. Un contains the unit(s) present in the new subset but not in the old one Un(1,2) for example contains the unit included in step init+1 Un(end,2) contains the units included in the final step of the search |
N |
Original contingency table, in array format. |
Ntable |
Original contingency table in table format (if initially supplied). |
Y |
array of size I-by-J containing row profiles. |
class |
'FSCorAnaeda' |
References
Atkinson, A.C., Riani, M. and Cerioli, A. (2004), "Exploring multivariate data with the forward search", Springer Verlag, New York.