MMregcore
MMregcore computes MM regression estimators for a selected fixed scale.
Description
Examples
MMregcore with all default options.
n=200;
p=3;
state1=123456;
randn('state', state1);
X=randn(n,p);
y=randn(n,1);
kk=10;
ycont = y;
ycont(1:kk)=ycont(1:kk)+7;
[outS]=Sreg(ycont,X);
outMM=MMregcore(ycont,X,outS.beta,outS.scale)
 MMregcore with optional input arguments.
MMregcore with optional input arguments.
 MMregcore with optional input arguments.
MMregcore with optional input arguments.Determine, e.g., an S estimate and extract the required arguments for the MM estimate.
n=200;
p=3;
state1=123456;
randn('state', state1);
X=randn(n,p);
y=randn(n,1);
kk=10;
ycont = y;
ycont(1:kk)=ycont(1:kk)+7;
[outS]=Sreg(ycont,X);
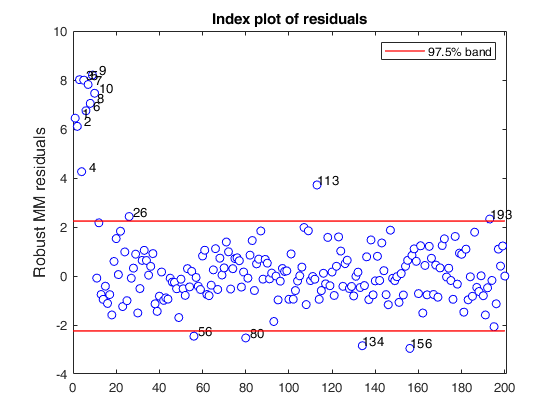
out=MMregcore(ycont,X,outS.beta,outS.scale,'plots',1)
Warning: Using 'state' to set RANDN's internal state causes RAND, RANDI, and
RANDN to use legacy random number generators. This syntax is not recommended.
See <a href="matlab:helpview([docroot '\techdoc\math\math.map'],'update_random_number_generator')">Replace Discouraged Syntaxes of rand and randn</a> to use RNG to replace the
old syntax.
Total estimated time to complete S estimate: 0.72 seconds
out =
struct with fields:
class: 'MMreg'
beta: [4×1 double]
weights: [200×1 double]
residuals: [200×1 double]
outliers: [1.00 2.00 3.00 4.00 5.00 6.00 7.00 8.00 … ] (1×17 double)
conflev: 0.97
rhofunc: 'bisquare'
rhofuncparam: []
Related Examples
Weighting the residuals with a rho function.
Determine, e.g., an S estimate and extract the required arguments for the MM estimate.
% This time use a Tukey biweight for S estimation and HA rho function
% for MM loop.
n=200;
p=3;
state1=123456;
randn('state', state1);
X=randn(n,p);
y=randn(n,1);
kk=10;
ycont = y;
ycont(1:kk)=ycont(1:kk)+7;
[outS]=Sreg(ycont,X);
rhofunc='hampel';
outMM1=MMregcore(ycont,X,outS.beta,outS.scale,'rhofunc',rhofunc,'plots',1)
Comparison between direct call to MMreg and call to Sreg and MMregcore.
In this example, two different rho functions are used for S and MM.
n=30;
p=3;
randn('state', 16);
X=randn(n,p);
% Uncontaminated data
y=randn(n,1);
% Contaminated data
ycont=y;
ycont(1:5)=ycont(1:5)+6;
% Two different rho functions are used for S and MM
rhofuncS='hyperbolic';
rhofuncMM='hampel';
% Direct call to MMreg
[out]=MMreg(ycont,X,'Srhofunc',rhofuncS,'rhofunc',rhofuncMM,'Snsamp',0);
% Call to Sreg and then to MMregcore
[outS]=Sreg(ycont,X,'rhofunc',rhofuncS,'nsamp',0);
outMM=MMregcore(ycont,X,outS.beta,outS.scale,'rhofunc',rhofuncMM);
disp('Difference between direct call to S and the calls to Sreg and MMregcore')
max(abs([out.beta-outMM.beta]))
Input Arguments
y — Response variable.
Vector.
A vector with n elements that contains the response variable. y can be either a row or a column vector.
Data Types: single| double
X — Data matrix of explanatory variables (also called 'regressors') of
dimension (n x p-1).
Rows of X represent observations, and columns
represent variables.
Missing values (NaN's) and infinite values (Inf's) are allowed, since observations (rows) with missing or infinite values will automatically be excluded from the computations.
Data Types: single| double
b0 — Initial estimate of beta.
Vector.
Vector containing initial estimate of beta (generally an S estimate with high breakdown point (e.g. 0.5).
Data Types: single| double
auxscale — scale estimate.
Scalar.
Scalar containing estimate of the scale (generally an S estimate with high breakdown point (e.g. .5)).
Data Types: single| double
Name-Value Pair Arguments
Specify optional comma-separated pairs of Name,Value arguments.
Name is the argument name and Value
is the corresponding value. Name must appear
inside single quotes (' ').
You can specify several name and value pair arguments in any order as
Name1,Value1,...,NameN,ValueN.
'intercept',false
, 'eff',0.99
, 'effshape',1
, 'refsteps',10
, 'tol',1e-10
, 'conflev',0.99
, 'rhofunc','optimal'
, 'rhofuncparam',5
, 'nocheck',true
, 'plots',0
, 'yxsave',1
intercept
—Indicator for constant term.true (default) | false.
Indicator for the constant term (intercept) in the fit, specified as the comma-separated pair consisting of 'Intercept' and either true to include or false to remove the constant term from the model.
Example: 'intercept',false
Data Types: boolean
eff
—nominal efficiency.scalar.
Scalar defining nominal efficiency (i.e. a number between 0.5 and 0.99). The default value is 0.95 Asymptotic nominal efficiency is:
Example: 'eff',0.99
Data Types: double
effshape
—location or scale efficiency.dummy scalar.
If effshape=1 efficiency refers to shape efficiency, else (default) efficiency refers to location.
Example: 'effshape',1
Data Types: double
refsteps
—Maximum iterations.scalar.
Scalar defining maximum number of iterations in the MM loop. Default value is 100.
Example: 'refsteps',10
Data Types: double
reftol
—Tolerance.scalar.
Scalar controlling tolerance in the MM loop.
Default value is 1e-7
Example: 'tol',1e-10
Data Types: double
conflev
—Confidence level which is
used to declare units as outliers.scalar.
Usually conflev=0.95, 0.975 0.99 (individual alpha) or 1-0.05/n, 1-0.025/n, 1-0.01/n (simultaneous alpha).
Default value is 0.975
Example: 'conflev',0.99
Data Types: double
rhofunc
—rho function.string.
String which specifies the rho function which must be used to weight the residuals.
Possible values are: 'bisquare';
'optimal';
'hyperbolic';
'hampel';
'mdpd'.
'AS'.
'bisquare' uses Tukey's \rho and \psi functions.
See TBrho and TBpsi.
'optimal' uses optimal \rho and \psi functions.
See OPTrho and OPTpsi.
'hyperbolic' uses hyperbolic \rho and \psi functions.
See HYPrho and HYPpsi.
'hampel' uses Hampel \rho and \psi functions.
See HArho and HApsi.
'mdpd' uses Minimum Density Power Divergence \rho and \psi functions.
'AS' uses Andrew's sine \rho and \psi functions.
The default is bisquare.
Example: 'rhofunc','optimal'
Data Types: char
rhofuncparam
—Additional parameters for the specified rho function.scalar | vector | empty value.
For hyperbolic rho function it is possible to set up the value of k = sup CVC (the default value of k is 4.5).
For Hampel rho function it is possible to define parameters a, b and c (the default values are a=2, b=4, c=8). For the other rho functions (Tuhey, PD and optimal), it is an empty value.
Example: 'rhofuncparam',5
Data Types: single | double
nocheck
—Check input arguments.boolean.
If nocheck is equal to true, no check is performed on matrix y and matrix X. Notice that y and X are left unchanged. In other words, the additional column of ones for the intercept is not added.
As default nocheck=false.
Example: 'nocheck',true
Data Types: boolean
plots
—Plot on the screen.scalar | structure.
If plots = 1, generates a plot with the robust residuals against index number. The confidence level used to draw the confidence bands for the residuals is given by the input option conflev. If conflev is not specified, a nominal 0.975 confidence interval will be used.
Example: 'plots',0
Data Types: single | double
yxsave
—the response vector y and data matrix X are saved into the output
structure out.scalar.
Default is 0, i.e. no saving is done.
Example: 'yxsave',1
Data Types: double
Output Arguments
out — description
Structure
A structure containing the following fields
| Value | Description |
|---|---|
beta |
p x 1 vector. Estimate of beta coefficients after refsteps refining steps. |
residuals |
n x 1 vector containing the estimates of the robust scaled residuals. |
outliers |
A vector containing the list of the units declared as outliers using confidence level specified in input scalar conflev. |
conflev |
Confidence level that was used to declare outliers. |
weights |
n x 1 vector. Weights assigned to each observation. |
rhofunc |
string identifying the rho function which has been used. |
rhofuncparam |
vector which contains the additional parameters for the specified rho function that have been used. For hyperbolic rho function the value of k =sup CVC. For Hampel rho function the parameters a, b and c. This field is empty if rhofunc is not 'hampel' or 'hyperbolic'. |
y |
response vector y. The field is present only if option yxsave is set to 1. |
X |
data matrix X. The field is present only if option yxsave is set to 1. |
class |
'MMreg' |
More About
Additional Details
It does iterative reweighted least squares (IRWLS) steps from "initial beta" (b0) keeping the estimate of the scale (auxscale) fixed.
References
Maronna, R.A., Martin D. and Yohai V.J. (2006), "Robust Statistics, Theory and Methods", Wiley, New York.
Acknowledgements
This function follows the lines of MATLAB/R code developed during the years by many authors.
For more details see the R library robustbase http://robustbase.r-forge.r-project.org/.
The core of these routines, e.g. the resampling approach, however, has been completely redesigned, with considerable increase of the computational performance.