FSRBr
Bayesian forward search in linear regression reweighted
Syntax
Description
FSRBr uses the units not declared as outliers by FSRB to produce a robust fit.
The units whose residuals exceeds the threshold determined by option alpha are declared as outliers. Moreover, it is possible in option R2th to modify the estimate of sigma2 which is used to declare the outliers. This is useful when there is almost a perfect fit in the data, the estimate of the error variance is very small and therefore there is the risk of declaring as outliers very small deviations from the robust fit. In this case, the estimate of sigma2 is corrected in order to achieve a value of R2 equal to R2th.
Example of FSRBr for international trade data (explore options).out
=FSRBr(y,
X,
Name, Value)
Examples
 Example of FSRBr for international trade data.
Example of FSRBr for international trade data.
 Example of FSRBr for international trade data.
Example of FSRBr for international trade data.Bayesian FS to fit the group of undervalued flows.
load('fishery');
X = fishery{:,1};
y = fishery{:,2};
[n,p] = size(X);
X = X + 0.000001*randn(n,1);
% id = undervalued flows
id = (y./X < 9.5);
% my prior on beta
mybeta = median(y(id)./X(id));
bmad = mad(y(id)./X(id));
ListIn = find(y./X <= mybeta + bmad);
% gscatter(X,y,id)
rr = y(ListIn)-(X(ListIn,:) * mybeta);
numS2cl = rr'*rr;
dfe = length(ListIn)-p;
S2cl = numS2cl/dfe;
bayes = struct;
bayes.R = X(ListIn,:)'*X(ListIn,:);
bayes.tau0 = 1/S2cl;
bayes.n0 = length(ListIn);
bayes.beta0 = mybeta;
% fit based on Bayesian FS with prior on the underdeclared flows
[out_B, xnew1 , ypred1, yci1] = FSRBr(y,X,'bayes',bayes,'intercept',false,'alpha',0.01,'bonflev',0.999,'fullreweight',false,'plotsPI',1,'plots',0);
h1 = allchild(gca); a1 = gca; f1 = gcf;
% fit based on traditional FS
[out, xnew2 , ypred2, yci2] = FSRr(y,X,'intercept',false,'alpha',0.01,'bonflev',0.999,'fullreweight',false,'plotsPI',1,'plots',0);
h2 = allchild(gca); a2 = gca; f2 = gcf;
% move the figure above into a single one with two panels
hh = figure; ax1 = subplot(2,1,1); ax2 = subplot(2,1,2);
copyobj(h1,ax1); title(ax1,get(get(a1,'title'),'string'));
copyobj(h2,ax2); title(ax2,get(get(a2,'title'),'string'));
figsize = get(hh, 'Position');
set(hh,'Position',figsize);
close(f1); close(f2);
disp(['Bayesian FS fit = ' num2str(out_B.betar) ' using a prior based on undervalued flows']);
disp(['Traditional FS fit = ' num2str(out.betar)]);Bayesian FS fit = 8.565 using a prior based on undervalued flows Traditional FS fit = 13.0939
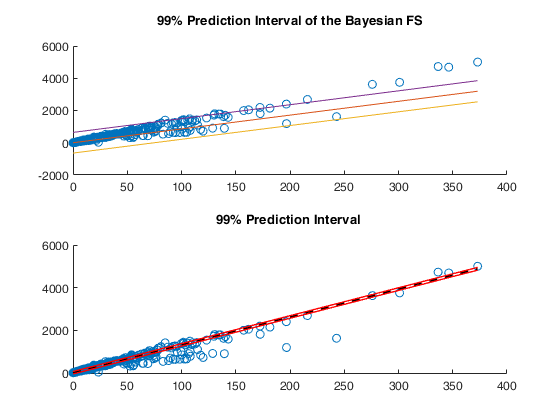
Example of FSRBr for international trade data (explore options).
Bayesian FS to fit the group of undervalued flows.
load('fishery');
X = fishery{:,1};
y = fishery{:,2};
[n,p] = size(X);
X = X + 0.000001*randn(n,1);
% id = undervalued flows
id = (y./X < 9.5);
% my prior on beta
mybeta = median(y(id)./X(id));
bmad = mad(y(id)./X(id));
ListIn = find(y./X <= mybeta + bmad);
% gscatter(X,y,id)
rr = y(ListIn)-(X(ListIn,:) * mybeta);
numS2cl = rr'*rr;
dfe = length(ListIn)-p;
S2cl = numS2cl/dfe;
bayes = struct;
bayes.R = X(ListIn,:)'*X(ListIn,:);
bayes.tau0 = 1/S2cl;
bayes.n0 = length(ListIn);
bayes.beta0 = mybeta;
alpha=0.0001;
bonflev=0.99999;
% fit based on Bayesian FS with prior on the underdeclared flows
[out_B, xnew1 , ypred1, yci1] = FSRBr(y,X,'bayes',bayes,'intercept',0,'alpha',alpha,'bonflev',bonflev,'fullreweight',false,'plotsPI',1,'plots',0);
h1 = allchild(gca); a1 = gca; f1 = gcf;
% fit based on traditional FS
[out, xnew2 , ypred2, yci2] = FSRr(y,X,'intercept',0,'alpha',alpha,'bonflev',bonflev,'fullreweight',false,'plotsPI',1,'plots',0);
h2 = allchild(gca); a2 = gca; f2 = gcf;
% move the figure above into a single one with two panels
hh = figure; ax1 = subplot(2,1,1); ax2 = subplot(2,1,2);
copyobj(h1,ax1); title(ax1,get(get(a1,'title'),'string'));
copyobj(h2,ax2); title(ax2,get(get(a2,'title'),'string'));
figsize = get(hh, 'Position');
set(hh,'Position',figsize);
close(f1); close(f2);
disp(['Bayesian FS fit = ' num2str(out_B.betar) ' using a prior based on undervalued flows']);
disp(['Traditional FS fit = ' num2str(out.betar)]);
Input Arguments
y — Response variable.
Vector.
Response variable, specified as a vector of length n, where n is the number of observations. Each entry in y is the response for the corresponding row of X.
Missing values (NaN's) and infinite values (Inf's) are allowed, since observations (rows) with missing or infinite values will automatically be excluded from the computations.
Data Types: single| double
X — Predictor variables.
Matrix.
Matrix of explanatory variables (also called 'regressors') of dimension n x (p-1) where p denotes the number of explanatory variables including the intercept.
Rows of X represent observations, and columns represent variables. By default, there is a constant term in the model, unless you explicitly remove it using input option intercept, so do not include a column of 1s in X. Missing values (NaN's) and infinite values (Inf's) are allowed, since observations (rows) with missing or infinite values will automatically be excluded from the computations.
Data Types: single| double
Name-Value Pair Arguments
Specify optional comma-separated pairs of Name,Value arguments.
Name is the argument name and Value
is the corresponding value. Name must appear
inside single quotes (' ').
You can specify several name and value pair arguments in any order as
Name1,Value1,...,NameN,ValueN.
'alpha',0.01
, 'R2th',0.95
, 'fullreweight',true
, 'plotsPI',1
, 'intercept',false
, bayes=struct;bayes.R=R;bayes.n0=n0;bayes.beta0=beta0;bayes.tau0=tau0;
, 'plots',1
, 'tag',{'plmdr' 'plyXplot'};
, 'init',100 starts monitoring from step m=100
, 'nocheck',true
, 'bivarfit',2
, 'multivarfit','1'
, 'labeladd','1'
, 'nameX',{'NameVar1','NameVar2'}
, 'namey','NameOfResponse'
, 'bonflev',0.99
, 'msg',1
alpha
—test size.scalar.
Number between 0 and 1 which defines test size to declare the outliers
Example: 'alpha',0.01
Data Types: double
R2th
—R2 threshold.scalar.
Scalar which defines the value R2 does have to exceed. For example, if R2 based on good observations is 0.92 and R2th is 0.90 the estimate of the variance of the residuals which is used to declare the outliers is adjusted in order to have a value of R2 which is equal to 0.90.
Example: 'R2th',0.95
Data Types: double
fullreweight
—Option to declare outliers.boolean.
If fullreweight is true (default option), the list of outliers refers to all the units whose residuals is above the threshold else if it is false the outliers are the observations which by procedure FSR had been declared outliers and have a residual greater than threshold
Example: 'fullreweight',true
Data Types: boolean
plotsPI
—Plot of prediction intervals.scalar.
If plotsPI =1 and the number of regressors (excluding the constant term) is equal 1, it is possible to see on the screen the yX scatter with superimposed the prediction intervals using a confidence level 1-alpha, else no plot is shown on the screen
Example: 'plotsPI',1
Data Types: double
intercept
—Indicator for constant term.true (default) | false.
Indicator for the constant term (intercept) in the fit, specified as the comma-separated pair consisting of 'Intercept' and either true to include or false to remove the constant term from the model.
Example: 'intercept',false
Data Types: boolean
bayes
—Prior information.structure.
It contains the following fields
| Value | Description |
|---|---|
beta0 |
p-times-1 vector containing prior mean of \beta |
R |
p-times-p positive definite matrix which can be interpreted as X0'X0 where X0 is a n0 x p matrix coming from previous experiments (assuming that the intercept is included in the model). The prior distribution of $\tau_0$ is a gamma distribution with parameters $a_0$ and $b_0$, that is \[ p(\tau_0) \propto \tau^{a_0-1} \exp (-b_0 \tau) \qquad E(\tau_0) = a_0/b_0 \] |
tau0 |
scalar. Prior estimate of \[ \tau=1/ \sigma^2 =a_0/b_0 \] |
n0 |
scalar. Sometimes it helps to think of the prior information as coming from n0 previous experiments. Therefore we assume that matrix X0 (which defines R), was made up of n0 observations. REMARK if structure bayes is not supplied the default values which are used are. beta0= zeros(p,1): Vector of zeros. R=eye(p): Identity matrix. tau0=1/1e+6: Very large value for the prior variance, that is a very small value for tau0. n0=1: just one prior observation. $\beta$ is assumed to have a normal distribution with mean $\beta_0$ and (conditional on $\tau_0$) covariance $(1/\tau_0) (X_0'X_0)^{-1}$. $\beta \sim N( \beta_0, (1/\tau_0) (X_0'X_0)^{-1} )$ |
Example: bayes=struct;bayes.R=R;bayes.n0=n0;bayes.beta0=beta0;bayes.tau0=tau0;
Data Types: double
plots
—Plot on the screen.scalar.
If plots=1 (default) the plot of minimum deletion residual with envelopes based on n observations and the scatterplot matrix with the outliers highlighted is produced.
If plots=2 the user can also monitor the intermediate plots based on envelope superimposition.
Else no plot is produced.
Example: 'plots',1
Data Types: double
tag
—tags to the plots which are created.character | cell array of characters.
This option enables to add a tag to the plots which are created. The default tag names are: fsr_mdrplot for the plot of mdr based on all the observations;
fsr_yXplot for the plot of y against each column of X with the outliers highlighted;
fsr_resuperplot for the plot of resuperimposed envelopes. The first plot with 4 panel of resuperimposed envelopes has tag fsr_resuperplot1, the second fsr_resuperplot2 ...
If tag is character or a cell of characters of length 1, it is possible to specify the tag for the plot of mdr based on all the observations;
If tag is a cell of length 2 it is possible to control both the tag for the plot of mdr based on all the observations and the tag for the yXplot with outliers highlighted.
If tag is a cell of length 3 the third element specifies the names of the plots of resuperimposed envelopes.
Example: 'tag',{'plmdr' 'plyXplot'};
Data Types: char or cell
init
—Search initialization.scalar.
scalar which specifies the initial subset size to start monitoring exceedances of minimum deletion residual, if init is not specified it set equal to: p+1, if the sample size is smaller than 40;
min(3*p+1,floor(0.5*(n+p+1))), otherwise.
Example: 'init',100 starts monitoring from step m=100
Data Types: double
nocheck
—Check input arguments.boolean.
If nocheck is equal to 1 no check is performed on matrix y and matrix X. Notice that y and X are left unchanged. In other words, the additional column of ones for the intercept is not added. As default nocheck=0.
Example: 'nocheck',true
Data Types: boolean
bivarfit
—Superimpose bivariate least square lines.character.
This option adds one or more least square lines, based on SIMPLE REGRESSION of y on Xi, to the plots of y|Xi.
bivarfit = '' is the default: no line is fitted.
bivarfit = '1' fits a single ols line to all points of each bivariate plot in the scatter matrix y|X.
bivarfit = '2' fits two ols lines: one to all points and another to the group of the genuine observations. The group of the potential outliers is not fitted.
bivarfit = '0' fits one ols line to each group. This is useful for the purpose of fitting mixtures of regression lines.
bivarfit = 'i1' or 'i2' or 'i3' etc.
fits an ols line to a specific group, the one with index 'i' equal to 1, 2, 3 etc. Again, useful in case of mixtures.
Example: 'bivarfit',2
Data Types: char
multivarfit
—Superimpose multivariate least square lines.character.
This option adds one or more least square lines, based on MULTIVARIATE REGRESSION of y on X, to the plots of y|Xi.
multivarfit = '' is the default: no line is fitted.
multivarfit = '1' fits a single ols line to all points of each bivariate plot in the scatter matrix y|X. The line added to the scatter plot y|Xi is avconst + Ci*Xi, where Ci is the coefficient of Xi in the multivariate regression and avconst is the effect of all the other explanatory variables different from Xi evaluated at their centroid (that is overline{y}'C)) multivarfit = '2' equal to multivarfit ='1' but this time we also add the line based on the group of unselected observations (i.e. the normal units).
Example: 'multivarfit','1'
Data Types: char
labeladd
—Add outlier labels in plot.character.
If this option is '1', we label the outliers with the unit row index in matrices X and y. The default value is labeladd='', i.e. no label is added.
Example: 'labeladd','1'
Data Types: char
nameX
—Add variable labels in plot.cell array of strings.
cell array of strings of length p containing the labels of the variables of the regression dataset. If it is empty (default) the sequence X1, ..., Xp will be created automatically
Example: 'nameX',{'NameVar1','NameVar2'}
Data Types: cell
namey
—Add response label.character.
character containing the label of the response
Example: 'namey','NameOfResponse'
Data Types: char
ylim
—Control y scale in plot.vector.
vector with two elements controlling minimum and maximum on the y axis. Default value is '' (automatic scale)
Example: 'ylim','[0,10]' sets the minimum value to 0 and the
max to 10 on the y axis
Data Types: double
xlim
—Control x scale in plot.vector.
vector with two elements controlling minimum and maximum on the x axis. Default value is '' (automatic scale)
Example: 'xlim','[0,10]' sets the minimum value to 0 and the
max to 10 on the x axis
Data Types: double
bonflev
—Signal to use to identify outliers.scalar.
option to be used if the distribution of the data is strongly non normal and, thus, the general signal detection rule based on consecutive exceedances cannot be used. In this case bonflev can be: - a scalar smaller than 1 which specifies the confidence level for a signal and a stopping rule based on the comparison of the minimum MD with a Bonferroni bound. For example, if bonflev=0.99 the procedure stops when the trajectory exceeds for the first time the 99% bonferroni bound.
- A scalar value greater than 1. In this case the procedure stops when the residual trajectory exceeds for the first time this value.
Default value is '', which means to rely on general rules based on consecutive exceedances.
Example: 'bonflev',0.99
Data Types: double
msg
—Level of output to display.scalar.
scalar which controls whether to display or not messages on the screen If msg==1 (default) messages are displayed on the screen about step in which signal took place and ....
else no message is displayed on the screen
Example: 'msg',1
Data Types: double
Output Arguments
out — description
Structure
Structure which contains the following fields
| Value | Description |
|---|---|
outliers |
k x 1 vector containing the list of the units declared outliers by procedure FSR or NaN if the sample is homogeneous |
beta |
p-by-1 vector containing the estimated regression parameter by procedure FSR |
outliersr |
k1 x 1 vector containing the list of the units declared outliers after the reweighting step or NaN if the sample is homogeneous |
betar |
p-by-1 vector containing the estimated regression parameter after the reweighting step |
rstud |
n-by-2 matrix. First column = studentized residuals Second column = p-values (computed using as reference distribution the Student t) |
class |
'FSRBr'. |
varargout —xnew : new points.
Vector
Vector with a number of new points where to evaluate the prediction interval. xnew is a vector.
ypred : values predicted by the fitted model on xnew. Vector of length(xnew) yci : Confidence intervals. A two-column matrix with each row providing one interval.
References
Chaloner, K. and Brant, R. (1988), A Bayesian Approach to Outlier Detection and Residual Analysis, "Biometrika", Vol. 75, pp. 651-659.
Riani, M., Corbellini, A. and Atkinson, A.C. (2018), Very Robust Bayesian Regression for Fraud Detection, "International Statistical Review", http://dx.doi.org/10.1111/insr.12247
Atkinson, A.C., Corbellini, A. and Riani, M., (2017), Robust Bayesian Regression with the Forward Search: Theory and Data Analysis, "Test", Vol. 26, pp. 869-886, https://doi.org/10.1007/s11749-017-0542-6